Prediction and validation of the structural features of Ov58GPCR, an immunogenic determinant of Onchocerca volvulus

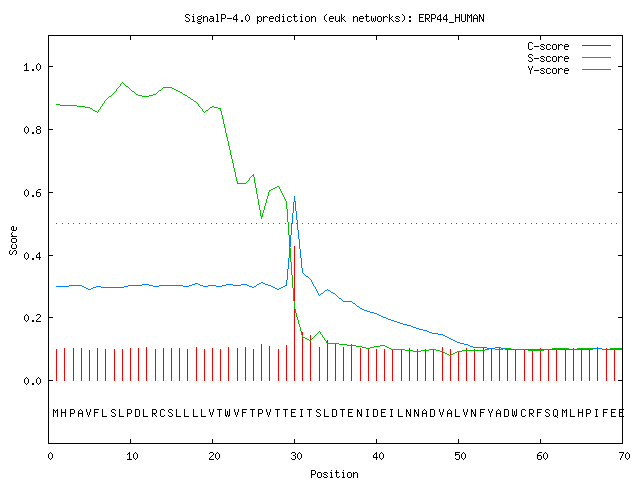
In silico analysis of the signal peptides sequences by SignalP 4.1 server. | Download Scientific Diagram

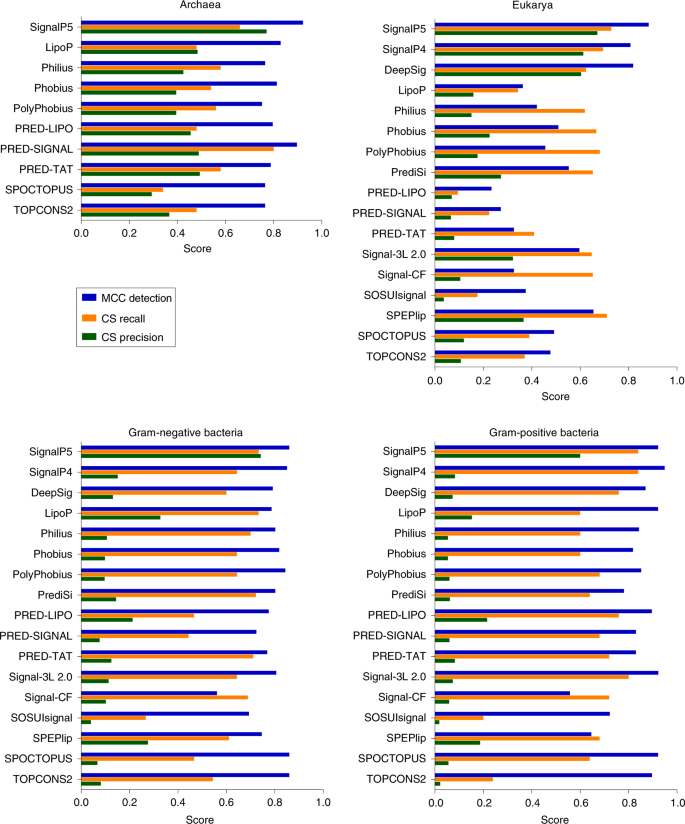
New tools for high‐throughput expression of fungal secretory proteins in Saccharomyces cerevisiae and Pichia pastoris - González - 2019 - Microbial Biotechnology - Wiley Online Library

β-Barrels covalently link peptidoglycan and the outer membrane in the α-proteobacterium Brucella abortus | Nature Microbiology
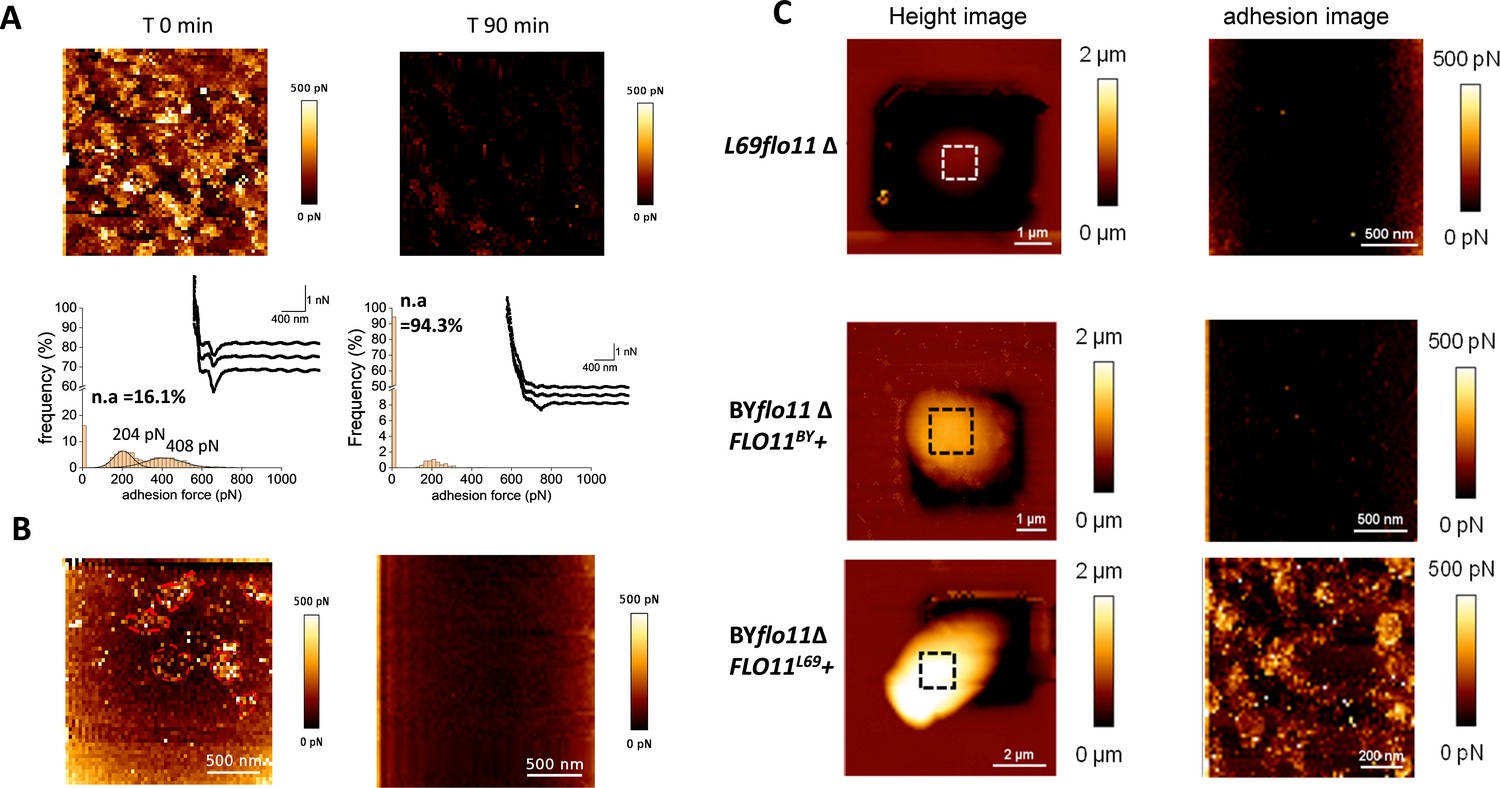
The dual role of amyloid-β-sheet sequences in the cell surface properties of FLO11-encoded flocculins in Saccharomyces cerevisiae | eLife
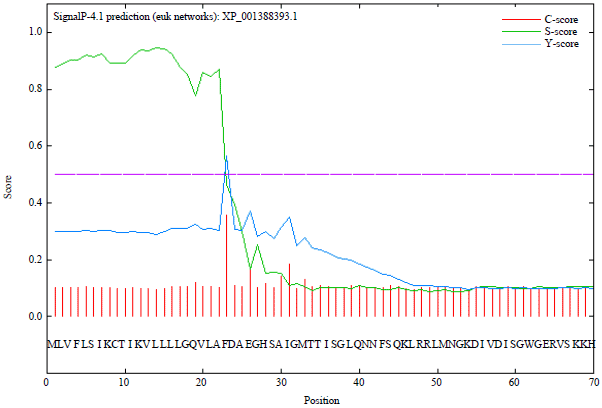
Signal Peptide Sequence Analysis of Selected Protein Sequences from Cryptosporidium parvum - SciAlert Responsive Version
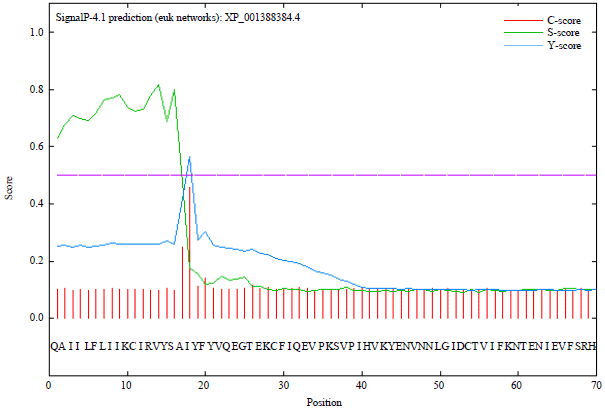
Signal Peptide Sequence Analysis of Selected Protein Sequences from Cryptosporidium parvum - SciAlert Responsive Version

In silico analysis of the signal peptides sequences by SignalP 4.1 server. | Download Scientific Diagram

BRAWNIN: A sORF-encoded Peptide Essential for Vertebrate Mitochondrial Complex III Assembly | bioRxiv
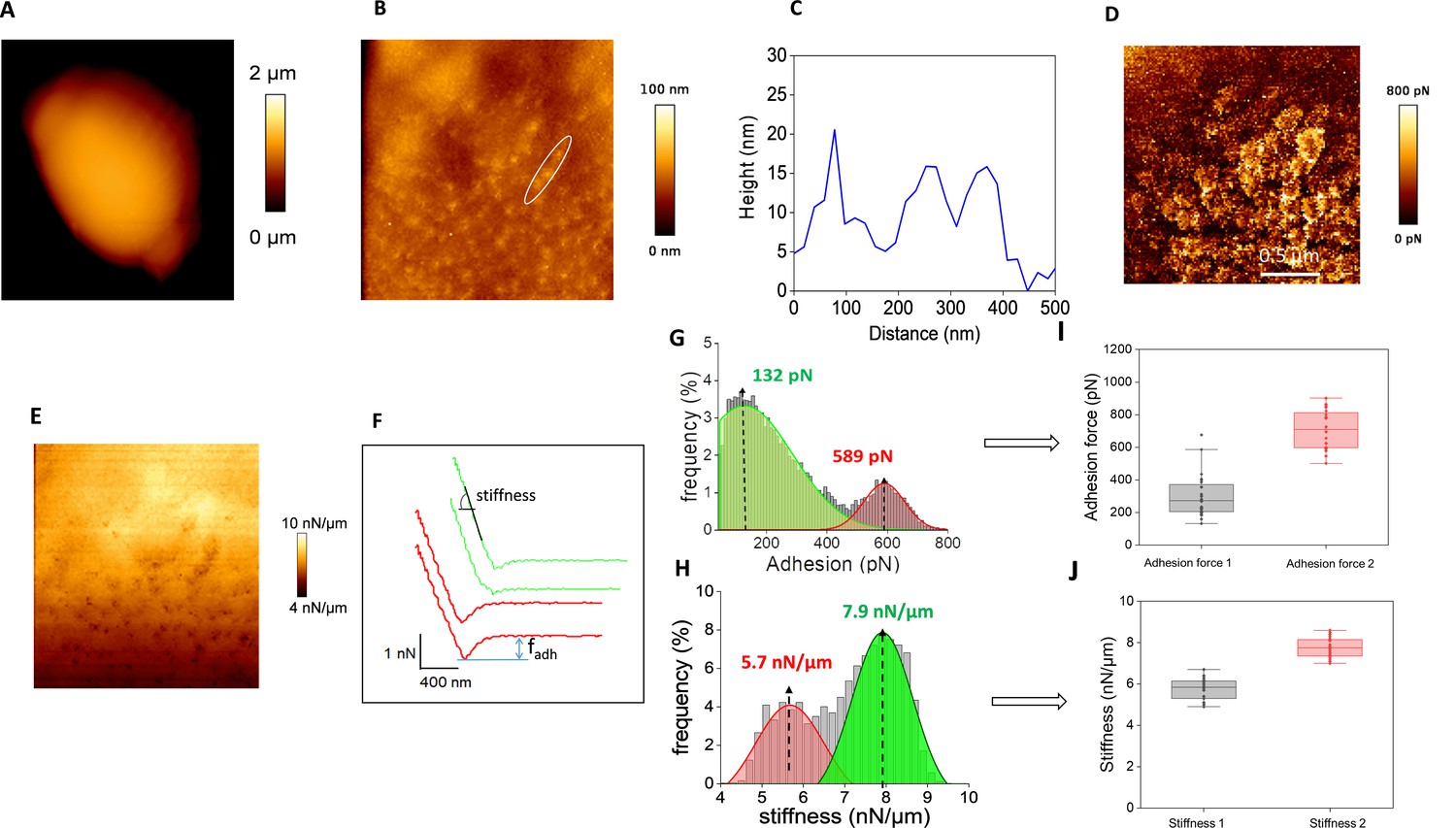
The dual role of amyloid-β-sheet sequences in the cell surface properties of FLO11-encoded flocculins in Saccharomyces cerevisiae | eLife

β-Barrels covalently link peptidoglycan and the outer membrane in the α-proteobacterium Brucella abortus | Nature Microbiology

New tools for high‐throughput expression of fungal secretory proteins in Saccharomyces cerevisiae and Pichia pastoris - González - 2019 - Microbial Biotechnology - Wiley Online Library

Microorganisms | Free Full-Text | Comparative Genomics of Marine Bacteria from a Historically Defined Plastic Biodegradation Consortium with the Capacity to Biodegrade Polyhydroxyalkanoates | HTML

β-Barrels covalently link peptidoglycan and the outer membrane in the α-proteobacterium Brucella abortus | Nature Microbiology
![Comparative genome analysis of 24 bovine-associated Staphylococcus isolates with special focus on the putative virulence genes [PeerJ] Comparative genome analysis of 24 bovine-associated Staphylococcus isolates with special focus on the putative virulence genes [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2018/4560/1/fig-2-full.png)
Comparative genome analysis of 24 bovine-associated Staphylococcus isolates with special focus on the putative virulence genes [PeerJ]

Sm29 signal peptide prediction using the SignalP 3.0 server according... | Download Scientific Diagram

SignalP 4.1 prediction of cellular localization and signal sequence... | Download Scientific Diagram